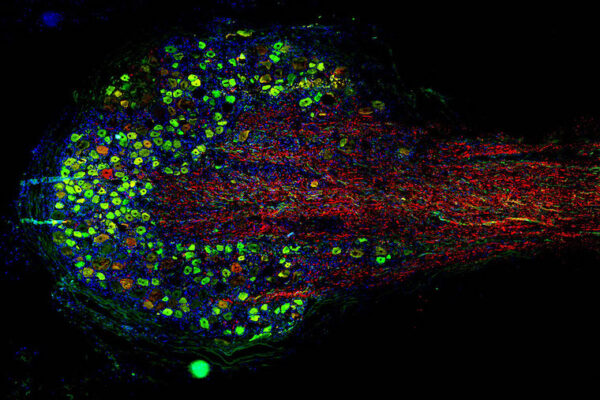
Researchers at WashU Medicine and collaborating institutions have developed a novel computational tool that can accurately identify a genetic problem in a gene called RFC1 that is linked to certain forms of peripheral neuropathy. Peripheral neuropathy is one of the most common neurological disorders and can cause pain, sensory loss, imbalance and weakness. It affects 12–20% of all people in the U.S. and can affect up to 30% of adults over age 65. The new research is published in Annals of Neurology.
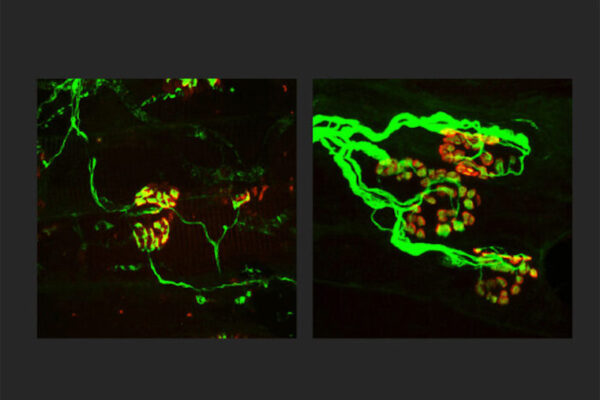
The disease-causing change, known as an RFC1 repeat expansion, has been associated with neuropathy, but its role across the broader spectrum of patients with unexplained, or “idiopathic,” neuropathy has remained unclear. One reason is that these repeat expansions — in which the set of DNA “letters” AAGGG is repeated many more times than normal — are difficult to detect using standard genetic testing methods.
The research team led by senior author Sheng Chih (Peter) Jin, an assistant professor of genetics and of pediatrics, and first author Zitian Tang, a graduate student in Jin’s lab, set out to bridge this technical gap by developing a new computational pipeline coupled with machine learning that can reliably identify and classify repeat expansions from genome sequencing data. Using this approach, they found that RFC1 repeat expansions may account for more than 2% of cases of idiopathic peripheral neuropathy.
The new tool offers a more affordable and reliable way to look for this extremely complex genetic variation in both clinical and research settings. The finding also supports broader genetic testing for people with unexplained peripheral neuropathy, including those who have muscle weakness as well as sensory symptoms. The team has made the tool public on GitHub, which could help expand testing to help more patients receive an accurate diagnosis and give families clearer information about the genetic causes of their condition.